Our mission
Accelerating time-to-Insights for personalized health care:
We empower individuals and organizations by providing near real-time health insights and analysis based on microbiome, proteome, epigenome and other biological data.
Our portfolio

Deepbiomics
This platform uses advanced graph and network-based representations, to create clear and simple summaries of complex biological processes, proteins, or gene sequences by breaking them into manageable components. It is a vital too

ElectricGut

OnSpotBiome

Proteome2Pathogen
As part of our ongoing commitment to transforming healthcare through near real-time biological insights we’re excited to introduce the Proteome2Pathogen Application. This advanced tool, developed in collaboration with

Symbaiose

MicroDiet
MicroDiet is more than just a nutrition app.It’s a sophisticated tool designed to explore the intricate relationship between dietary habits, gut microbiota, and metabolic disorders such as obesity and type 2 diabetes (T2D) across
Where microbiome meets diagnostics: MicroGnostics
HORAIZON's MicroGnostics platform embodies the principle of personalized medicine, recognizing the unique microbial fingerprint of each individual. Our approach not only enhances the accuracy of disease diagnosis but also allows for the customization of treatment strategies to suit the individual's microbiome composition.
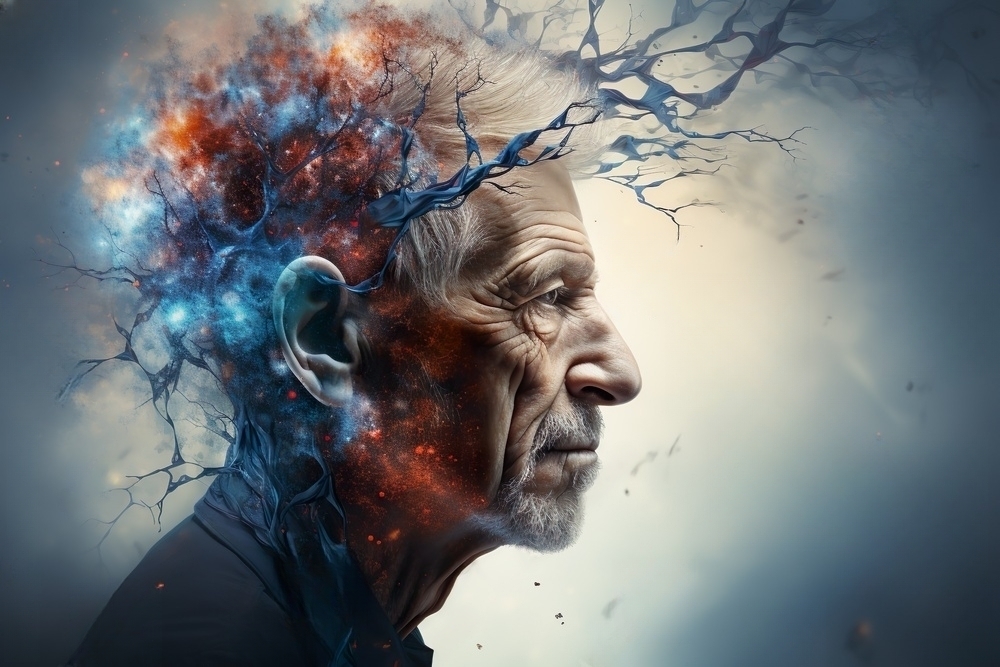

NeuraBiome
Linking mental health with microbiome and optimal nutrition
NeuraBiome stands at the forefront of this exciting field, leveraging advanced algorithms and machine learning techniques to analyze and interpret complex data. This non-invasive and efficient method surpasses traditional diagnostic tools in speed and accessibility and offers a more comprehensive understanding of the microbiome’s impact on neural health.
Our products for:
Personalized treatment plans

Business optimization

Time-to-insight
About us
We are a multidisciplinary team composed of AI specialists and life science researchers. Together, we create cutting-edge, interpretable AI models and algorithms tailored for diverse scientific domains, such as microbiome, epigenome and proteome. Our mission is to expedite the identification of new biomarkers and therapeutic targets, thereby revolutionising the diagnosis and treatment of a cardiometabolic diseases such as diabetes, Inflammatory bowel disease, cancer, and vascular disorders.

- 1. Data-powered solutions
- 2. Data complexity simplified
- 3. Enter HORAIZON
- 4. Empower decision-making with real-time insights
01. Data-powered solutions
02. Data complexity simplified
03. Enter HORAIZON
04. Empower decision-making with real-time insights
Expertise
Building
success through collaboration

Solutions

Impact

Innovation

Building customer relationships
Our clients, collaborators, EU partners














So, if you’re ready to unlock unparalleled value from your data, HORAIZON is your go-to partner for smarter, faster, and more effective data analytics.







